Frontiers of Data and Computing ›› 2024, Vol. 6 ›› Issue (3): 3-14.
CSTR: 32002.14.jfdc.CN10-1649/TP.2024.03.001
doi: 10.11871/jfdc.issn.2096-742X.2024.03.001
• Special Issue: Advance of Intelligent Healthcare • Previous Articles Next Articles
CAI Chengfei1,3( ),LI Jun2,JIAO Yiping2,WANG Xiangxue2,GUO Guanchen1,XU Jun2,*(
),LI Jun2,JIAO Yiping2,WANG Xiangxue2,GUO Guanchen1,XU Jun2,*( )
)
Received:2023-10-21
Online:2024-06-20
Published:2024-06-21
CAI Chengfei, LI Jun, JIAO Yiping, WANG Xiangxue, GUO Guanchen, XU Jun. Progress and Challenges of Medical Multimodal Data Fusion Methods Based on Deep Learning in Oncology[J]. Frontiers of Data and Computing, 2024, 6(3): 3-14, https://cstr.cn/32002.14.jfdc.CN10-1649/TP.2024.03.001.
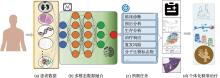
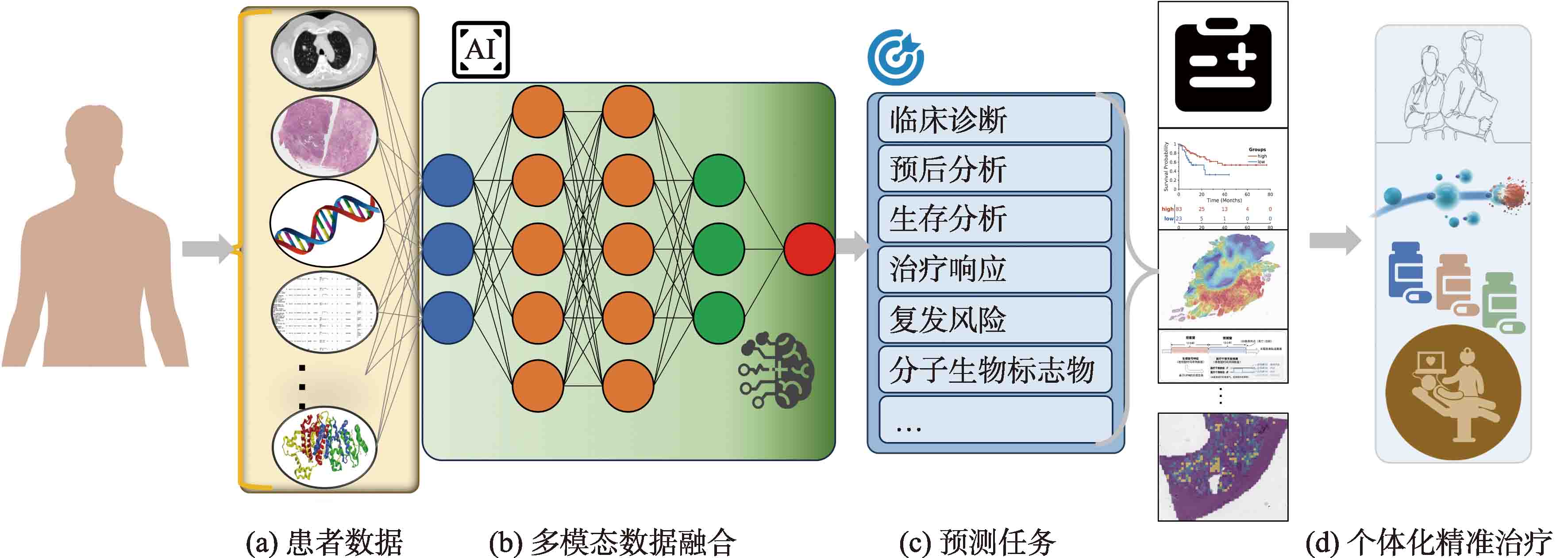
Fig.1
Presents an integrated framework that combines information from various data sources and clinical backgrounds, unfolding different prediction tasks to empower individualized precision healthcare (a) Patient datas; (b) Multimodal data fusion; (c) Prediction tasks; (d) Personalized precision treatment"
| [1] | GUO Z, et al. Medical image segmentation based on multi-modal convolutional neural network: Study on image fusion schemes[C]. 2018 IEEE 15th International Symposium on Biomedical Imaging, IEEE, 2018: 903-907. |
| [2] |
LI L, et al. Deep learning for variational multimodality tumor segmentation in PET/CT[J]. Neurocomputing, 2020, 392: 277-295.
doi: 10.1016/j.neucom.2018.10.099 pmid: 32773965 |
| [3] | SAEED N, et al. An ensemble approach for patient prognosis of head and neck tumor using multimodal data[J]. 3D Head and Neck Tumor Segmentation in PET/CT Challenge, Cham: Springer International Publishing, 2021, 13209: 278-286. |
| [4] | SHOEIBI A, et al. Diagnosis of brain diseases in fusion of neuroimaging modalities using deep learning: A review[J]. Information Fusion, 2022, 93: 85-117. |
| [5] | BERA K, et al. Artificial intelligence in digital pathology—new tools for diagnosis and precision oncology[J]. Nature reviews Clinical oncology, 2019, 16(11): 703-715. |
| [6] | VU Q D, et al. Handcrafted Histological Transformer (H2T): Unsupervised representation of whole slide images[J]. Medical Image Analysis, 2023. 85: 102743-102761. |
| [7] | HUANG Y, et al. Deep learning radiopathomics based on preoperative US images and biopsy whole slide images can distinguish between luminal and non-luminal tumors in early-stage breast cancers[J]. EBioMedicine, 2023, 94: 104706-104719. |
| [8] | BILAL M, et al. Development and validation of a weakly supervised deep learning framework to predict the status of molecular pathways and key mutations in colorectal cancer from routine histology images: a retrospective study[J]. The Lancet Digital Health, 2021, 3(12): 763-772. |
| [9] | YAMASHITA R, et al. Deep learning model for the prediction of microsatellite instability in colorectal cancer: a diagnostic study[J]. The Lancet Oncology, 2021, 22(1): 132-141. |
| [10] | CHANG X, et al. Predicting colorectal cancer microsatellite instability with a self-attention-enabled convolutional neural network[J]. Cell Reports Medicine, 2023, 4(2): 100914-100930. |
| [11] |
MARCUS L, et al. FDA approval summary: pembrolizumab for the treatment of microsatellite instability-high solid tumors[J]. Clinical Cancer Research, 2019, 25(13): 3753-3758.
doi: 10.1158/1078-0432.CCR-18-4070 pmid: 30787022 |
| [12] | LIU J, et al. Immune landscape and prognostic immune-related genes in KRAS-mutant colorectal cancer patients[J]. Journal of translational medicine, 2021, 19: 1-17. |
| [13] |
WAGNER S J, et al. Transformer-based biomarker prediction from colorectal cancer histology: A large-scale multicentric study[J]. Cancer Cell, 2023, 41(9): 1650-1661.
doi: 10.1016/j.ccell.2023.08.002 pmid: 37652006 |
| [14] | PAI R K, et al. Quantitative pathologic analysis of digitized images of colorectal carcinoma improves prediction of recurrence-free survival[J]. Gastroenterology, 2022, 163(6): 1531-1546. |
| [15] | LI Z, et al. Survival Prediction via Hierarchical Multimodal Co-Attention Transformer: A Computational Histology-Radiology Solution[J]. IEEE Transactions on Medical Imaging, 2023, 42(9): 2678-2689. |
| [16] | BOEHM K.M., et al. Harnessing multimodal data integration to advance precision oncology[J]. Nature Reviews Cancer, 2022, 22(2): 114-126. |
| [17] | RAZZAGHI P, et al. Multimodal brain tumor detection using multimodal deep transfer learning[J]. Applied Soft Computing, 2022, 129: 109631-109642. |
| [18] | XU L, et al. Bi-MGAN: Bidirectional T1-to-T2 MRI images prediction using multi-generative multi-adversarial nets[J]. Biomedical Signal Processing and Control, 2022, 78: 103994-104005. |
| [19] |
HAVAEI M, et al. Brain tumor segmentation with deep neural networks[J]. Medical image analysis, 2017, 35: 18-31.
doi: S1361-8415(16)30033-0 pmid: 27310171 |
| [20] | LI Y, et al. Deep learning based multimodal brain tumor diagnosis[C]. Springer International Publishing, 2018, Revised Selected Papers 3: 149-158. |
| [21] | GAO, ZEYU, et al. A semi-supervised multi-task learning framework for cancer classification with weak annotation in whole-slide images[J]. Medical Image Analysis, 2023, 83: 102652-102666. |
| [22] | GAO, ZEYU, et al. Uncertainty-based Model Acceleration for Cancer Classification in Whole-Slide Images[C]. 2022 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), IEEE, 2022, 1534-1538. |
| [23] | GRAHAM S, et al. Screening of normal endoscopic large bowel biopsies with artificial intelligence: a retrospective study[J]. Gut, 2023, 72: 1709-1721. |
| [24] | ZHANG S, et al. Graph convolutional networks: a comprehensive review[J]. Computational Social Networks, 2019, 6(1): 1-23. |
| [25] | QU L, et al. Towards label-efficient automatic diagnosis and analysis: a comprehensive survey of advanced deep learning-based weakly-supervised, semi-supervised and self-supervised techniques in histopathological image analysis[J]. Physics in Medicine and Biology, 2022, 67(20): 1-35. |
| [26] | FENG J and ZHOU Z. Deep miml network[C]. In Proceedings of the AAAI Conference on Artificial Intelligence (AAAI), 2017, 31(1): 10890-10897. |
| [27] | WANG X, et al. Revisiting multiple instance neural networks[J]. Pattern Recognition, 2018, 74: 15-24. |
| [28] | WU J, et al. Deep multiple instance learning for image classification and auto-annotation[C]. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2015: 3460-3469. |
| [29] | CHEN T, et al. A simple framework for contrastive learning of visual representations[C]. In International Conference on Machine Learning (ICML), 2020, 1597-1607. |
| [30] | KATHRER J N, et al. Multi-class texture analysis in colorectal cancer histology[J]. Scientific Reports, 2016, 6(1): 1-11. |
| [31] | HASHIMOTO N, et al. Multi-scale domain-adversarial multiple-instance cnn for cancer subtype classification with unannotated histopathological images[C]. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2020: 3852-3861. |
| [32] | LI H, et al. Dt-mil: Deformable transformer for multi-instance learning on histopathological image[C]. In International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI), Springer, 2021: 206-216. |
| [33] | YAO J, et al. Whole slide images based cancer survival prediction using attention guided deep multiple instance learning networks[J]. Medical Image Analysis, 2020, 65: 101789-101804. |
| [34] | ZHU X, et al. Wsisa: Making survival prediction from whole slide histopathological images[C]. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2017: 7234-7242. |
| [35] |
CAMPANELLA G, et al. Clinical-grade computational pathology using weakly supervised deep learning on whole slide images[J]. Nature medicine, 2019, 25(8): 1301-1309.
doi: 10.1038/s41591-019-0508-1 pmid: 31308507 |
| [36] |
LU M Y, et al. Data-efficient and weakly supervised computational pathology on whole-slide images[J]. Nature biomedical engineering, 2021, 5(6): 555-570.
doi: 10.1038/s41551-020-00682-w pmid: 33649564 |
| [37] | ILSE M, et al. Attention-based deep multiple instance learning[C]. International conference on machine learning, PMLR, 2018: 2127-2136. |
| [38] | DOSOVITSKIY A, et al. An image is worth 16x16 words: Transformers for image recognition at scale[J]. arXiv preprint, arXiv, 2020, 2010: 11929-11951. |
| [39] | VASWANI A, et al. Attention is all you need[C]. Advances in neural information processing systems, 2017: 30-41. |
| [40] | HU, ZEBIN, et al. Data-enabled intelligence in complex industrial systems cross-model transformer method for medical image synthesis[J]. Complexity, 2021, (2021): 1-7. |
| [41] | TRAGAKIS A, et al. The fully convolutional transformer for medical image segmentation[C]. Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision, 2023: 3660-3669. |
| [42] | JING L, et al. Self-supervised visual feature learning with deep neural networks: A survey[J]. IEEE transactions on pattern analysis and machine intelligence, 2020, 43(11): 4037-4058. |
| [43] | KAZEROUNI, et al. Diffusion models for medical image analysis: A comprehensive survey[J]. arXiv preprint arXiv:2211.07804 (2022). |
| [44] | SLEEMANIV W C, et al. Multimodal classification: Current landscape, taxonomy and future directions[J]. ACM Computing Surveys, 2022, 55(7): 1-31. |
| [45] | BALTRUSAITIS T, et al. Multimodal machine learning: A survey and taxonomy[J]. IEEE transactions on pattern analysis and machine intelligence, 2018, 41(2): 423-443. |
| [46] |
QIAN X, et al. Prospective assessment of breast cancer risk from multimodal multiview ultrasound images via clinically applicable deep learning[J]. Nature biomedical engineering, 2021, 5(6): 522-532.
doi: 10.1038/s41551-021-00711-2 pmid: 33875840 |
| [47] |
LE M.H., et al. Automated diagnosis of prostate cancer in multi-parametric MRI based on multimodal convolutional neural networks[J]. Physics in Medicine and Biology, 2017, 62(16): 6497-6514.
doi: 10.1088/1361-6560/aa7731 pmid: 28582269 |
| [48] |
LIPKOVA J, et al. Personalized radiotherapy design for glioblastoma: Integrating mathematical tumor models, multimodal scans, and bayesian inference[J]. IEEE transactions on medical imaging, 2019, 38(8): 1875-1884.
doi: 10.1109/TMI.2019.2902044 pmid: 30835219 |
| [49] |
NIE D, et al. Multi-channel 3D deep feature learning for survival time prediction of brain tumor patients using multi-modal neuroimages[J]. Scientific reports, 2019, 9(1): 1103-1117.
doi: 10.1038/s41598-018-37387-9 pmid: 30705340 |
| [50] |
YAP J, et al. Multimodal skin lesion classification using deep learning[J]. Experimental dermatology, 2018, 27(11): 1261-1267.
doi: 10.1111/exd.13777 pmid: 30187575 |
| [51] | XU T, et al. Multimodal deep learning for cervical dysplasia diagnosis[C]. Medical Image Computing and Computer-Assisted Intervention-MICCAI 2016: 19th, Springer International Publishing, 2016, Proceedings, Part II 19: 115-123. |
| [52] |
KHOSRAVI P, et al. A deep learning approach to diagnostic classification of prostate cancer using pathology-radiology fusion[J]. Journal of Magnetic Resonance Imaging, 2021, 54(2): 462-471.
doi: 10.1002/jmri.27599 pmid: 33719168 |
| [53] |
CHEN R J, et al. Pan-cancer integrative histology-genomic analysis via multimodal deep learning[J]. Cancer Cell, 2022, 40(8): 865-878.
doi: 10.1016/j.ccell.2022.07.004 pmid: 35944502 |
| [54] | SAMMUT S.J., et al. Multi-omic machine learning predictor of breast cancer therapy response[J]. Nature, 2022, 601(7894): 623-629. |
| [55] | RAMANATHAN T T, et al. Naïve Bayes Based Multiple Parallel Fuzzy Reasoning Method for Medical Diagnosis[J]. Journal of Engineering Science and Technology, 2022, 17(1): 472-490. |
| [56] | REDA I, et al. Deep learning role in early diagnosis of prostate cancer[J]. Technology in cancer research and treatment, 2018, 17: 1-11. |
| [57] | LU M Y, et al. AI-based pathology predicts origins for cancers of unknown primary[J]. Nature, 2021, 594(7861): 106-110. |
| [58] | SHA X, et al. Identifying pathological subtypes of non-small-cell lung cancer by using the radiomic features of 18F-fluorodeoxyglucose positron emission computed tomography[J]. Translational Cancer Research, 2019, 8(5): 1741-1749. |
| [59] |
JOO S, et al. Multimodal deep learning models for the prediction of pathologic response to neoadjuvant chemotherapy in breast cancer[J]. Scientific reports, 2021, 11(1): 18800-18808.
doi: 10.1038/s41598-021-98408-8 pmid: 34552163 |
| [60] | KUMAR A, et al. Co-learning feature fusion maps from PET-CT images of lung cancer[J]. IEEE Transactions on Medical Imaging, 2019, 39(1): 204-217. |
| [61] |
SEDGHI A, et al. Improving detection of prostate cancer foci via information fusion of MRI and temporal enhanced ultrasound[J]. International journal of computer assisted radiology and surgery, 2020, 15: 1215-1223.
doi: 10.1007/s11548-020-02172-5 pmid: 32372384 |
| [62] | HAVAEI M, et al. Hemis: Hetero-modal image segmentation[C]. Medical Image Computing and Computer-Assisted Intervention-MICCAI 2016: 19th, Springer International Publishing, 2016, Proceedings, Part II 19: 469-477. |
| [63] | LIANG M, et al. Integrative data analysis of multi-platform cancer data with a multimodal deep learning approach[J]. IEEE/ACM transactions on computational biology and bioinformatics, 2014, 12(4): 928-937. |
| [64] | LAI Y H, et al. Overall survival prediction of non-small cell lung cancer by integrating microarray and clinical data with deep learning[J]. Scientific reports, 2020, 10(1): 4679-4690. |
| [65] | VALE-SILVA L A, et al. Long-term cancer survival prediction using multimodal deep learning[J]. Scientific Reports, 2021, 11(1): 13505-13516. |
| [66] |
YALA A, et al. A deep learning mammography-based model for improved breast cancer risk prediction[J]. Radiology, 2019, 292(1): 60-66.
doi: 10.1148/radiol.2019182716 pmid: 31063083 |
| [67] | MO S, et al. Multimodal priors guided segmentation of liver lesions in MRI using mutual information based graph co-attention networks[C]. Medical Image Computing and Computer Assisted Intervention-MICCAI 2020: 23rd, Springer International Publishing, 2020, Proceedings, Part IV 23:429-438. |
| [68] | LEI B, et al. Self-co-attention neural network for anatomy segmentation in whole breast ultrasound[J]. Medical image analysis, 2020, 64: 101753-101769. |
| [69] | ZHOU L and YAN L. Deep features fusion with mutual attention transformer for skin lesion diagnosis[C]. 2021 IEEE International Conference on Image Processing (ICIP), IEEE, 2021: 3797-3801. |
| [70] | VO H Q, et al. Multimodal Breast Lesion Classification Using Cross-Attention Deep Networks[C]. 2021 IEEE EMBS International Conference on Biomedical and Health Informatics (BHI), IEEE, 2021: 1-4. |
| [1] | LIAO Libo, WANG Shudong, SONG Weimin, ZHANG Zhaoling, LI Gang, HUANG Yongsheng. The Study of Jet Tagging Algorithm Based on DeepSets at CEPC [J]. Frontiers of Data and Computing, 2024, 6(3): 108-115. |
| [2] | YAN Jin, DONG Kejun, LI Hongtao. A Deep Web Tracker Detection Method with Coordinated Semantic and Co-Occurrence Features [J]. Frontiers of Data and Computing, 2024, 6(3): 127-138. |
| [3] | WANG Zhiyong, LIU Jingjing, WANG Xinming, CHEN Bowen, NIE Wei, ZHANG Hanlin, LIU Honghai. Advancements and Frontiers in Autism Diagnosis and Treatment Based on Artificial Intelligence [J]. Frontiers of Data and Computing, 2024, 6(3): 15-27. |
| [4] | KOU Dazhi. Automatic Teeth Segmentation on Dental Panoramic Radiographs with Deep Learning [J]. Frontiers of Data and Computing, 2024, 6(3): 162-172. |
| [5] | ZHENG Yinuo, SUN Muyi, ZHANG Hongyun, ZHANG Jing, DENG Tianzheng, LIU Qian. Application of Deep Learning in Dental Implant Imaging: Research Progress and Challenges [J]. Frontiers of Data and Computing, 2024, 6(3): 41-49. |
| [6] | YUAN Jialin, OUYANG Rushan, DAI Yi, LAI Xiaohui, MA Jie, GONG Jingshan. Assessing the Clinical Utility of a Deep Learning-Based Model for Calcification Recognition and Classification in Mammograms [J]. Frontiers of Data and Computing, 2024, 6(2): 68-79. |
| [7] | WANG Ziyuan, WANG Guozhong. Application of Improved Lightweight YOLOv5 Algorithm in Pedestrian Detection [J]. Frontiers of Data and Computing, 2023, 5(6): 161-172. |
| [8] | JU Jiaji, HUANG Bo, ZHANG Shuai, GUO Ruyan. A Dual-Channel Sentiment Analysis Model Integrating Sentiment Lexcion and Self-Attention [J]. Frontiers of Data and Computing, 2023, 5(4): 101-111. |
| [9] | LI JunFei, XU LiMing, WANG Yang, WEI Xin. Review of Automatic Citation Classification Based on Deep Learning Technology [J]. Frontiers of Data and Computing, 2023, 5(4): 86-100. |
| [10] | LI Yan,HE Hongbo,WANG Runqiang. A Survey of Research on Microblog Popularity Prediction [J]. Frontiers of Data and Computing, 2023, 5(2): 119-135. |
| [11] | LIU Yunfan,LI Qi,SUN Zhenan,TAN Tieniu. Face Age Editing Methods Based on Generative Adversarial Network: A Survey [J]. Frontiers of Data and Computing, 2023, 5(2): 2-23. |
| [12] | TU Youyou,ZHENG Qijing,ZHAO Jin. Research on Quantum Proton Coupled Charge Transfer Process Based on Deep Neural Network [J]. Frontiers of Data and Computing, 2023, 5(2): 37-49. |
| [13] | XU Songyuan,LIU Feng. ESDRec: A Data Recommendation Model for Earth Big Data Platform [J]. Frontiers of Data and Computing, 2023, 5(1): 55-64. |
| [14] | LIU Qiwei,LI Jun,GU Beibei,ZHAO Zefang. TSAIE: Text Sentiment Analysis Model Based on Image Enhancement [J]. Frontiers of Data and Computing, 2022, 4(3): 131-140. |
| [15] | CHEN Qiong,YANG Yong,HUANG Tianlin,FENG Yuan. A Survey on Few-Shot Image Semantic Segmentation [J]. Frontiers of Data and Computing, 2021, 3(6): 17-34. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
